
Molecular aspects of aneuploidy in preimplantation human embryos: a mini-review
Introduction
Overwhelmingly, aneuploidy has been observed to be the most significant obstacle in achieving successful pregnancies in in vitro fertilization (IVF) therapy. Specifically, more than 50% of preimplantation embryos are aneuploid and unable to achieve pregnancies resulting in live births (1). By definition, aneuploidy refers to acquisition of chromosomal copy numbers that differ from euploid chromosomal content. Aneuploidy has been observed in multiple embryo categories, irrespective of the mother’s age, and across many morphological classifications including arrested and developing embryos, fragmented embryos and even good morphology embryos in both fresh and frozen-thawed cycles (2-4). Aneuploidy in sex chromosomes is compatible with life and has mild effects in mental disability and growth alterations, although is known to cause infertility (5). This is due to X chromosome silencing through an epigenetic mediated pathway, in addition to the small genetic contribution of the Y chromosome (6). While some autosomal forms of trisomy do occur and contribute to significant birth effects including Patau syndrome (trisomy 13), Edwards syndrome (trisomy 18) and Down syndrome (trisomy 21) (7), the vast majority of autosomal aneuploidies are not compatible with life. In a follow-up study correlating the aneuploidy mechanisms with types of infertility, it was found that the repeated implantation failure group had the lowest proportion of meiotic errors (8.9%), for an average maternal age of 36 years, whereas the recurrent miscarriage group had the highest rate (24%), for an average maternal age of 37.6 years. The embryos from patients with repeated implantation failure were more prone to post-zygotic errors, specifically chaotic type complex errors (8). In human preimplantation embryos there are two types of aneuploidy; meiotic aneuploidy and mitotic aneuploidy (mosaicism). In this mini review we focus and analyze on the two aneuploidy types, their causes and the underlying molecular mechanisms involved.
Meiotic origin aneuploidy
Analysis of human sperm and oocytes has showed that aneuploidy in preimplantation embryos is mainly caused by an error-prone meiotic chromosome segregation mechanism in oocytes. In reproductive aged women, aneuploidy rate in human oocytes reaches 20–30%, in contrast to human sperm, in which only 1–8% have an abnormal chromosomal content (9,10). Trisomy 21, the most frequently occurring viable aneuploidy in humans, is caused due to missegregation of chromosomes 21 in female meiosis I (MI) (65%) or in meiosis II (MII) (23%) in contrast to trisomy 16 (which is incompatible with life) that in close to 100% of cases is due to missegregation in female MI (11-13). Maternal age is the major critical factor related to aneuploidy; 50% of the oocytes from advanced age women (≥40 years old) are aneuploid due to meiotic errors. In contrast, the incidence of aneuploidy in sperm is independent of paternal age (9,10). At least 5% of men diagnosed with infertility are at high risk of producing aneuploid sperm due to a major chromosomal abnormality. However, the cell cycle checkpoints during meiosis ensure that most potentially aneuploidy gametes undergo apoptotic cell death, leading to a lowered sperm count (14). In male gametes, most aneuploidies occur in the sex chromosomes because XY chromosomes maintain only a limited region of homology in order to pair and separate, in contrary to female meiosis where XX chromosome pair harbour multiple regions of homology.
In oocytes, there are three possible routes by which aneuploidy, specifically trisomy, can occur during meiotic divisions: (I) the ‘MI’, ‘homologue’, ‘true’ or ‘classic’ non-disjunction; in this case, during MI an extra chromosome arises through the lack of segregation of a bivalent between the oocyte and first polar body; (II) the MII non-disjunction; in this case, during MII the lack of segregation of the sister chromatid pair between the oocyte and the second polar body leads to trisomy; and (III) the MI or MII pre-division; in this case, at some point in MI or MII, before or during the segregation event, the normal pairing of chromosomes may not fulfilled. In MI, this could lead to the generation of two pairs of sister chromatids, called univalent, formed by the breaking down of a bivalent. Similarly, in MII, the dyad could be prematurely resolved into two single chromatids (15) (Figure 1).
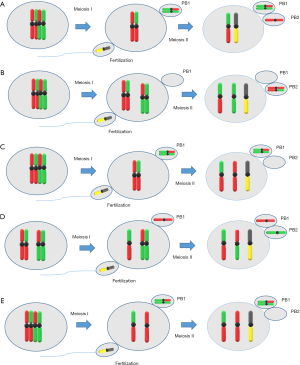
Meiosis is a specialized cell division process that creates genetically distinct haploid cells after the DNA replication step following two cell divisions (16). The newly replicated sister chromatids are connected by cohesin, a protein which maintains cohesion between sister chromatids at the first cell division (17). In this mechanism, seperase, a cysteine protease, is responsible for cleaving kleisin, a cohesin component (18). Furthermore, on resumption of MI, the dissociation of homologous chromosomes is initiated by the cleavage of cohesin subunit Rec8 by the seperase, along the chromosome arms (19). After that, sister chromatids are separated during MII, triggered by the cleavage of centromeric cohesin complex. Errors in this process may result in either chromosome loss or gain, depending on whether chromatids/homologs segregate to the polar body or to the oocyte, respectively. Despite that, premature separation of sister chromatids and reverse segregation, during the dictyate stage, involve loss of cohesion between sister chromatids possibly due to deterioration of cohesin protein complex (20).
It is known that the cohesive connections between chromosomes are weakened as maternal age advances (13,21). Specifically, cohesin is mainly loaded onto newly replicated chromosomes in oogonia, during fetal life (22). Despite that, oocytes have only a limited capacity to reload the cohesin complex once S phase is complete (22) and consequently cohesin will be reduced over time (23,24). Indeed, on bivalents from aged oocytes that still retain their integrity, very small amounts of Rec8 were detected (25). Similarly, in a study performed in dictyate stage human oocytes, it was highlighted an age-related decrease in cohesin subunits, Rec8 and SMC1β (26). Cohesin loss in aged oocytes can modify the proximity of the two sister kinetochores, a proteinaceous structure located at the centromeric DNA of each sister chromatid, affecting dramatically the efficiency of that pair to establish an attachment to just one pole. As a result, if sister kinetochores start acting independently and not as a pair, incorrect attachment to microtubules will be enhanced resulting in incorrect anaphase segregation (15).
The classic mechanism for meiotic aneuploidy is non-disjunction of whole chromosome or sister chromatids, in both MI and MII (27). However, data from karyotyping of human metaphase II oocytes from IVF patients, showed that at least some forms of trisomy resulted from malsegregation of single chromatids in MI, possibly from premature pre-division of the whole chromosomes into sister chromatids, which then segregate randomly (28). Another significant factor responsible for meiotic trisomies, besides advanced maternal age, is the reduced number or altered distribution of recombination events between chromosomes. Specifically, recombination can be interrupted in three different stages during the development of the mature oocyte: (I) prenatally with factors influencing recombination patterns before arrest in prophase of MI; (II) during follicle recruitment and growth by age-related changes; and (III) before ovulation and fertilization (which may occur several decades later) during resumption and completion of meiosis (29).
In summary, there is not a unique non-disjunctional mechanism applies for all chromosomes, as the mechanisms of non-disjunction and the influence of advanced maternal age among chromosomes vary significantly. However, these differences are not simply dependent on chromosome size, as might be expected if the loss of cohesin proteins was the only implicated mechanism. Rather, altered recombination and maternal age relationship is totally dependent on chromosomal context. This can be explained from the fact that recombination failure has been linked to some forms of trisomy (30,31) involving older women, but at the same time a chromosome-specific relationship between altered location of crossovers and age is also reported (32). Age-related loss of cohesion between homologous chromosomes is a possible mechanism for some situations of meiotic aneuploidy like in small size chromosomes held together by a single crossover, but it is not adequate in the case where chromosomes are held together by multiple and/or proximal crossovers. Thus, the evidence from human studies indicates that there are multiple mechanisms accompanying the maternal age effect.
Mitotic origin aneuploidy (mosaicism)
Genomic errors can also arise during post-fertilization mitotic divisions, resulting in embryonic mosaicism. By definition, embryonic mosaicism describes the presence of two or more chromosomally distinct populations of cells within the same embryo. Embryonic mosaicism is related with genetic diseases, miscarriages and preimplantation embryo loss (11).
As mitosis begins, the microtubules that are attached on the kinetochores of each chromosome begin to depolymerize, effectively trafficking chromosomes towards the opposite poles of the spindle. At the moment that sister chromatids are aligned on opposite poles, cytokinesis initiates and the cell divides, forming two identical cells (33). In contrast to some mammals, first mitotic division in human preimplantation embryos does not hold mitotic spindle checkpoints in order to prevent the cell from unequal chromosome divisions (34). For this reason, chromosome malsegregation and subsequent mitotic aneuploidy occurs in preimplantation human embryos (35). The different mechanisms leading to embryonic mosaicism during embryonic mitosis are: non-disjunction, anaphase lagging and endoreplication (Figure 2).
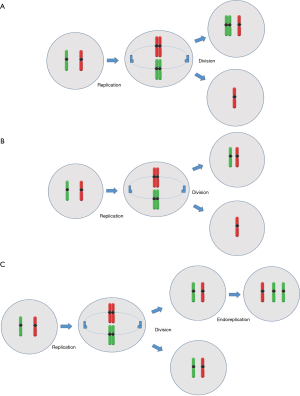
Non-disjunction is the failure of sister chromatids to separate correctly from each other during mitosis, resulting in a cell with a monosomy and another cell with a trisomy. The timing during which non-disjunction occurs is very important for the type of mosaicism. If it occurs prior to cell differentiation, such as in cleavage stage embryos (<8 cells), then a general mosaicism is created. If it occurs after differentiation in the trophoblast cells, then only the placenta cells will be mosaic while the actual embryo will be euploid. The latest is called confined mosaicism because it is present only in one particular area (e.g., placenta) (33). Non-disjunction is established as the main malsegragation mechanism in sex chromosomes during the first cleavage stage mitotic division (36), but is rarely associated with meiotic aneuploidy in autosome chromosomes (27,37). From the above one may conclude that chromosomes may be more or less susceptible to non-disjunction depending on the development stage. In addition, although embryonic mosaicism in general appears largely independent of maternal age, analysis of a large number of cleavage stage embryos showed a significant association between mitotic aneuploidy due to non-disjunction and maternal age (38).
Anaphase lagging during mitosis is the failure of a sister chromatid to properly separate from the other one because of improper spindle formation during anaphase, resulting in monosomy of that chromosome in one cell and disomy in the same chromosome in the other cell. The two main reasons behind anaphase lagging are: the failure of the chromatid to attach to the spindle or the failure of the attached to the spindle chromatid to be incorporated to the nucleus. Like in non-disjunction, the timing that anaphase lagging occurs determines the type of mosaicism (general mosaic or confined mosaicism) (33). In a study that was performed on discarded Day 5–6 embryos, it was found that monosomy occurred seven times more than trisomy (39), implicating anaphase lagging as the main source of mitotic aneuploidy in human preimplantation embryos.
Studies show that lagging chromosomes follow the common degradation pathway, they are often encapsulated in small, nucleus-like bodies separated from the main nucleus, the micronuclei (40). These chromosomes are correlated to severe DNA damage that can possibly impair their ability to create functional kinetochores and separate properly (41). Double-strand breaks, commonly observed in cleavage-stage embryos, contributing to mosaic patterns of segmental rearrangements (e.g., translocations) (42). Segmental replication of chromosomal followed by end-to-end fusion, can result in chromosome carrying two centromeres. These types of chromosomes are susceptible to breakage-fusion-bridge cycles, leading to a cascade of complex rearrangement in descendent cells (43). Furthermore, in subsequent mitotic divisions, the micronuclei may fuse with neighboring blastomeres or be reabsorbed into the main nucleus, resulting in complex mosaicism patterns (40). Micronuclei may also be expelled from the blastomeres by cellular fragmentation in which their chromosomes are degraded (44). Additionally, in a study correlating morphologic nuclear abnormalities with mosaicism in human preimplantation embryos, it was found that embryos containing blastomeres with abnormal micronuclei were aneuploid due to mitotic mosaicism, while embryos without abnormal nuclei were euploid or had only a low percentage of mosaicism. Specifically, blastomeres in embryos with micronucleation showed reciprocal chromosome gains and losses of both whole chromosome and of chromosome segments. Therefore, extra-nuclear DNA formation may possibly be a primary mechanism of mitotic aneuploidy (45).
Endoreplication defines the replication of a chromosome during the S phase of the cell cycle without the subsequent completion of mitosis, resulting in trisomic chromosome in one cell and a disomic chromosome in the other. Endoreplication can occur by two mechanisms: by a malfunction in cell cycle resulting to a replicated chromosome without subsequent cytokinesis or from a sudden shutdown after mitosis initiation that leads in a replicated chromosome (33). The vast amount of chromosomal aneuploidies within the human embryo is due to non-disjunction and anaphase lagging, while the incidence of endoreplication occur to a lesser extent (3).
The frequency of mitotic aneuploidies in human preimplantation embryos can be calculated from the mosaic embryos frequency. One relative systematic review and meta-analysis of studies showed that 73% of all human preimplantation embryos after IVF were mosaic. Interestingly, 22% of those embryos were diploid mosaic and 5% of them were detected with other abnormalities (46). Among mosaic embryos, the most common type was diploid-aneuploid embryos (59% of all embryos) followed by aneuploid mosaic embryos (15% of all embryos). The observed heterogeneity in the reported frequency of mosaic embryos is dependent on several factors including: (I) the number of chromosomes that have been analyzed; (II) the type of embryos (fresh vs. frozen); (III) the definition of mosaicism that is used; (IV) the developmental stage of the embryos (cleavage stage vs. blastocyst stage); and (V) the method of analysis that is used in a study.
In human preimplantation embryos, mitotic aneuploidies and mosaicism are very common during the first cleavage divisions. Specifically, ~15–90% of all cleavage stage human embryos are mosaic (47). In addition, meta-analysis data showed that mitotic aneuploidies increased from 63% at the cleavage stage to 95% to blastocyst stage (46). A relative study reported that 69% of abnormal blastocysts from advanced age women were mosaic for both the inner mass and the trophoectoderm (48). Even though the mosaic embryos percentage is higher at the blastocyst stage, the proportion of aneuploidy cells within the embryo decreases, resulting in relatively more diploid cells in blastocyst stage embryos (74%) compared to cleavage stage embryos (62%) (46,49-52).
At mitosis there are specific cell cycle checkpoints that ensure correct cell division before progression to the next stage. Relaxation of those cell cycle checkpoints can lead to ploidy mosaicism by occasionally allowing cells to skip M phase. This results to DNA replication without cell division, creating a tetraploid cell. Mosaic tetraploidy is very common to occur at earlier embryonic stages but can also be observed in molar pregnancies (<1%) (53). Moreover, when tetraploidy is accompanied by amplification of centrosome it can lead to multipolar cell division in following mitoses (44). The cell cycle checkpoints at mitosis occur during G1, G2 and metaphase stages of the cell cycle (54).
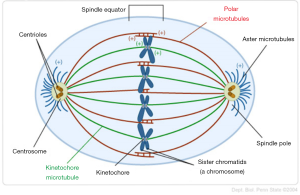
The G1 checkpoint controls all the required factors for DNA synthesis, such as quantity of energy, environment, cell size, and presence of nucleotides and nutrients. The G2 checkpoint controls proper DNA duplication during the S phase of the cell cycle and progression to mitosis. A relevant study reported that retinoblastoma protein, the key protein of the G1 cell cycle checkpoint, and Wee1-like protein kinase, the key protein of G2 cell cycle checkpoint, are absent from morphologically normal appearing Day-3 human embryos (55). In addition to the aforementioned cell cycle checkpoints, the primary checkpoint ensuring proper chromosome segregation is the spindle assembly checkpoint (SAC). As it is known, mitotic spindle consists of microtubules emerging from the spindle poles (Figure 3). Defects in microtubule dynamics, in spindle pole function or in kinetochore composition contribute to increased frequency of chromosome segregation errors. Components of SAC, such as Mad1, Mad2, Mad3, Mps1, Bub1, Bub3, BuBR1 and CENP-E recognize kinetochores that are empty or incorrectly attached and they activate cell cycle delay until the correct attachment of all chromosomes to microtubules and the proper alignment at the metaphase plate (56). This delay is performed via inhibition of the anaphase-promoting complex/cyclosome, whose activity is required for the metaphase-to-anaphase progression (57). Defects in SAC can lead to aneuploidy, both in vitro and in vivo, due to increased chromosome missegregation levels. Relaxation or absence of cell cycle checkpoints in early human preimplantation embryos contribute to aneuploidy by allowing a blastomere with chromosomal defects to enter and continue to mitotic divisions (58,59).
In addition to SAC, sister chromatid cohesion is vital for the maintenance of structural integrity of chromosomes and for proper attachment of chromosomes in the mitotic spindle (60). Cohesins function to keep the two sister chromatids connected, preventing them from premature separation from S-phase until anaphase. At anaphase, the association between the sister chromatids is ended allowing them separate each other. Cohesins consist of one stromal antigen (STAG) subunit (STAG1, STAG2, STAG3), two structural maintenance of chromosome (SMC) subunits (SMC1α, SMC1β, SMC3) and one kleisin subunit (RAD21, RAD21L, Rec8) (61). Most of the subunits are shared in both meiosis and mitosis, except subunits SMC1β, STAG3, RAD21L and Rec8 that are specific only to meiosis (61). Malfunction of cohesins leads in premature separation of chromosomes, while delay in their removal may lead to non-disjunction (62). Specifically, inactivation of STAG2 was found to lead in aneuploidy in human cells (63). The rate of mitotic errors in somatic cells was found to be increased by mutations in cohesin subunits and their regulators. Specifically, it was found that overexpression of seperase can frequently cause premature separation of sister chromatids, leading to aneuploidies involving both chromosome losses and gains (64). Similarly, knockout of several cohesin genes (including PLK1, STAG1, RAD21, NIPBL and SMC3) in mouse models resulted in early embryonic arrest (65). Furthermore, zygotes predicted to give rise to arrested embryos exhibit differential expression of several cohesin genes including SMC3 and securin, a protein involved in control of the metaphase-anaphase transition and anaphase onset (66). Finally, paternal factors are also related to mitotic aneuploidy, since severe sperm defects can increase the percentage of mitotic abnormalities and chaotic mosaic embryos. Specifically, embryos originating from patients with non-obstructive azoospermia, where sperm was retrieved after testicular sperm extraction, have an increased mosaicism rate compared to embryos derived from ejaculated sperm (53% and 26.5% respectively) (67) Furthermore, advanced paternal age (≥50 years old) induces mitotic aneuploidy and decreases blastocyst rate formation in embryos originated from donated oocytes (68).
Conclusions
The observed high rate of chromosomal aneuploidies in preimplantation human embryos may arise mostly during the first or second meiotic divisions, especially in advanced maternal age, but also can arise in the postzygotic stage during the mitotic divisions of cleavage stage and blastocyst embryos, resulting in mosaicism. The major etiological factor of meiotic aneuploidy is the maternal age. Cohesive connections holding together chromosomes are weakened with increasing maternal age possibly due to very small amounts of a major cohesin component, Rec8. This alternates the proximity of the two sister kinetochores promoting incorrect attachment to microtubules that finally will result in incorrect segregation in anaphase. Another significant correlation of meiotic aneuploidy (especially in trisomies) is with the reduced number or altered distribution of recombination events in various chromosomes. Intriguingly, the relationship between maternal age and altered recombination is entirely dependent on the context of chromosomes (Figure 4A). The summation of evidence from human studies clearly indicates that multiple mechanisms synergize with maternal age effect to promote aneuploidy.
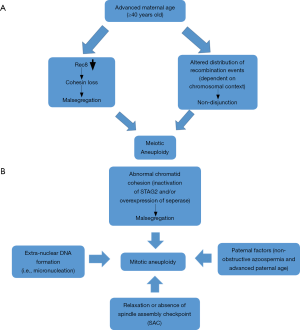
In contrast, mitotic aneuploidy (mosaicism) occurs more specifically, mainly by the relaxation or absence of cell cycle checkpoints via SAC which leads to aneuploidy both in vitro and in vivo arising from increased chromosome missegregation levels. In addition to SAC, sister chromatid cohesion is vital for the maintenance of structural integrity of chromosomes and for proper attachment of chromosomes in the mitotic spindle. Thus, malfunction of cohesins, possibly due to inactivation of STAG2 and/or overexpression of seperase, leads in premature separation of chromosomes, while delay in their removal may lead to non-disjunction. In addition, extra-nuclear DNA formation (micronucleation) from lagging chromosomes with severe DNA damage, may possibly be a primary mechanism of mitotic aneuploidy. Paternal factor is also related to mitotic aneuploidy, since advanced paternal age and severe sperm defects can increase the percentage of mitotic abnormalities and chaotic mosaic embryos, respectively (Figure 4B).
To move the field forward, we suggest further work and analysis to be performed in the molecular interactions between Rec8 protein with other cohesins (i.e., STAG3, SMC1β and RAD21L) in order to reveal the effects of advanced maternal age on sister chromatid cohesion throughout the whole meiotic process in human oocytes. Similarly, further studies can be possibly performed in the gene expression levels of different SAC components (i.e., Mad1, Mad2, Mad3, Mps1, Bub1, Bub3, BuBR1 and CENP-E) in order to elucidate the correlation between advanced maternal age and the inability of the SAC to detect certain attachment errors to chromosome kinetochores in the mitotic spindle.
Acknowledgements
The authors wish to acknowledge their appreciation to Dr. Larry L. Luchsinger for his valuable assistance in editing the manuscript’s grammar and structure. In addition, the authors would like to thank Prof. Richard Cyr for kindly granting his permission in order to include his figure (the 3rd illustration) in our study.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Franasiak JM, Forman EJ, Hong KH, et al. The nature of aneuploidy with increasing age of the female partner: a review of 15,169 consecutive trophectoderm biopsies evaluated with comprehensive chromosomal screening. Fertil Steril 2014;101:656-63.e1. [Crossref] [PubMed]
- Baart EB, Martini E, van den Berg I, et al. Preimplantation genetic screening reveals a high incidence of aneuploidy and mosaicism in embryos from young women undergoing IVF. Hum Reprod 2006;21:223-33. [Crossref] [PubMed]
- Daphnis DD, Fragouli E, Economou K, et al. Analysis of the evolution of chromosome abnormalities in human embryos from Day 3 to 5 using CGH and FISH. Mol Hum Reprod 2008;14:117-25. [Crossref] [PubMed]
- Santos MA, Teklenburg G, Macklon NS, et al. The fate of the mosaic embryo: chromosomal constitution and development of Day 4, 5 and 8 human embryos. Hum Reprod 2010;25:1916-26. [Crossref] [PubMed]
- Lenroot RK, Lee NR, Giedd JN. Effects of sex chromosome aneuploidies on brain development: evidence from neuroimaging studies. Dev Disabil Res Rev 2009;15:318-27. [Crossref] [PubMed]
- Prestel M, Feller C, Becker PB. Dosage compensation and the global re-balancing of aneuploid genomes. Genome Biol 2010;11:216. [Crossref] [PubMed]
- Hassold T, Hall H, Hunt P. The origin of human aneuploidy: where we have been, where we are going. Hum Mol Genet 2007;16 Spec No. 2:R203-8.
- Mantzouratou A, Mania A, Fragouli E, et al. Variable aneuploidy mechanisms in embryos from couples with poor reproductive histories undergoing preimplantation genetic screening. Hum Reprod 2007;22:1844-53. [Crossref] [PubMed]
- Wang J, Fan HC, Behr B, et al. Genome-wide single-cell analysis of recombination activity and de novo mutation rates in human sperm. Cell 2012;150:402-12. [Crossref] [PubMed]
- Lu S, Zong C, Fan W, et al. Probing meiotic recombination and aneuploidy of single sperm cells by whole-genome sequencing. Science 2012;338:1627-30. [Crossref] [PubMed]
- Hassold T, Hunt P. To err (meiotically) is human: the genesis of human aneuploidy. Nat Rev Genet 2001;2:280-91. [Crossref] [PubMed]
- Hassold T, Abruzzo M, Adkins K, et al. Human aneuploidy: incidence, origin, and etiology. Environ Mol Mutagen 1996;28:167-75. [Crossref] [PubMed]
- Nagaoka SI, Hassold TJ, Hunt PA. Human aneuploidy: mechanisms and new insights into an age-old problem. Nat Rev Genet 2012;13:493-504. [Crossref] [PubMed]
- Miharu N. Chromosome abnormalities in sperm from infertile men with normal somatic karyotypes: oligozoospermia. Cytogenet Genome Res 2005;111:347-51. [Crossref] [PubMed]
- Jones KT, Lane SI. Molecular causes of aneuploidy in mammalian eggs. Development 2013;140:3719-30. [Crossref] [PubMed]
- Page SL, Hawley RS. The genetics and molecular biology of the synaptonemal complex. Annu Rev Cell Dev Biol 2004;20:525-58. [Crossref] [PubMed]
- Petronczki M, Siomos MF, Nasmyth K. Un menage a quatre: the molecular biology of chromosome segregation in meiosis. Cell 2003;112:423-40. [Crossref] [PubMed]
- Nasmyth K. Cohesin: a catenase with separate entry and exit gates? Nat Cell Biol 2011;13:1170-7. [Crossref] [PubMed]
- Webster A, Schuh M. Mechanisms of Aneuploidy in Human Eggs. Trends Cell Biol 2017;27:55-68. [Crossref] [PubMed]
- Ottolini CS, Newnham L, Capalbo A, et al. Genome-wide maps of recombination and chromosome segregation in human oocytes and embryos show selection for maternal recombination rates. Nat Genet 2015;47:727-35. [Crossref] [PubMed]
- Jessberger R. Age-related aneuploidy through cohesion exhaustion. EMBO Rep 2012;13:539-46. [Crossref] [PubMed]
- Revenkova E, Herrmann K, Adelfalk C, et al. Oocyte cohesin expression restricted to predictyate stages provides full fertility and prevents aneuploidy. Curr Biol 2010;20:1529-33. [Crossref] [PubMed]
- Duncan FE, Hornick JE, Lampson MA, et al. Chromosome cohesion decreases in human eggs with advanced maternal age. Aging Cell 2012;11:1121-4. [Crossref] [PubMed]
- Sakakibara Y, Hashimoto S, Nakaoka Y, et al. Bivalent separation into univalents precedes age-related meiosis I errors in oocytes. Nat Commun 2015;6:7550. [Crossref] [PubMed]
- Chiang T, Duncan FE, Schindler K, et al. Evidence that weakened centromere cohesion is a leading cause of age-related aneuploidy in oocytes. Curr Biol 2010;20:1522-8. [Crossref] [PubMed]
- Tsutsumi M, Fujiwara R, Nishizawa H, et al. Age-related decrease of meiotic cohesins in human oocytes. PLoS One 2014;9:e96710. [Crossref] [PubMed]
- Handyside AH, Montag M, Magli MC, et al. Multiple meiotic errors caused by predivision of chromatids in women of advanced maternal age undergoing in vitro fertilisation. Eur J Hum Genet 2012;20:742-7. [Crossref] [PubMed]
- Angell RR. Predivision in human oocytes at meiosis I: a mechanism for trisomy formation in man. Hum Genet 1991;86:383-7. [Crossref] [PubMed]
- Hassold T, Hunt P. Maternal age and chromosomally abnormal pregnancies: what we know and what we wish we knew. Curr Opin Pediatr 2009;21:703-8. [Crossref] [PubMed]
- Fisher JM, Harvey JF, Morton NE, et al. Trisomy 18: studies of the parent and cell division of origin and the effect of aberrant recombination on nondisjunction. Am J Hum Genet 1995;56:669-75. [PubMed]
- Bugge M, Collins A, Petersen MB, et al. Non-disjunction of chromosome 18. Hum Mol Genet 1998;7:661-9. [Crossref] [PubMed]
- Oliver TR, Feingold E, Yu K, et al. New insights into human nondisjunction of chromosome 21 in oocytes. PLoS Genet 2008;4:e1000033. [Crossref] [PubMed]
- Taylor TH, Gitlin SA, Patrick JL, et al. The origin, mechanisms, incidence and clinical consequences of chromosomal mosaicism in humans. Hum Reprod Update 2014;20:571-81. [Crossref] [PubMed]
- Homer HA, McDougall A, Levasseur M, et al. Mad2 prevents aneuploidy and premature proteolysis of cyclin B and securin during meiosis I in mouse oocytes. Genes Dev 2005;19:202-7. [Crossref] [PubMed]
- Braude P, Bolton V, Moore S. Human gene expression first occurs between the four- and eight-cell stages of preimplantation development. Nature 1988;332:459-61. [Crossref] [PubMed]
- Bean CJ, Hassold TJ, Judis L, et al. Fertilization in vitro increases non-disjunction during early cleavage divisions in a mouse model system. Hum Reprod 2002;17:2362-7. [Crossref] [PubMed]
- Forman EJ, Treff NR, Stevens JM, et al. Embryos whose polar bodies contain isolated reciprocal chromosome aneuploidy are almost always euploid. Hum Reprod 2013;28:502-8. [Crossref] [PubMed]
- Munné S, Sandalinas M, Escudero T, et al. Chromosome mosaicism in cleavage-stage human embryos: evidence of a maternal age effect. Reprod Biomed Online 2002;4:223-32. [Crossref] [PubMed]
- Ioannou D, Fonseka KG, Meershoek EJ, et al. Twenty-four chromosome FISH in human IVF embryos reveals patterns of post-zygotic chromosome segregation and nuclear organisation. Chromosome Res 2012;20:447-60. [Crossref] [PubMed]
- Chavez SL, Loewke KE, Han J, et al. Dynamic blastomere behaviour reflects human embryo ploidy by the four-cell stage. Nat Commun 2012;3:1251. [Crossref] [PubMed]
- Vázquez-Diez C, Yamagata K, Trivedi S, et al. Micronucleus formation causes perpetual unilateral chromosome inheritance in mouse embryos. Proc Natl Acad Sci U S A 2016;113:626-31. [Crossref] [PubMed]
- Vanneste E, Voet T, Le Caignec C, et al. Chromosome instability is common in human cleavage-stage embryos. Nat Med 2009;15:577-83. [Crossref] [PubMed]
- Voet T, Vanneste E, Van der Aa N, et al. Breakage-fusion-bridge cycles leading to inv dup del occur in human cleavage stage embryos. Hum Mutat 2011;32:783-93. [Crossref] [PubMed]
- McCoy RC. Mosaicism in Preimplantation Human Embryos: When Chromosomal Abnormalities Are the Norm. Trends Genet 2017;33:448-63. [Crossref] [PubMed]
- Kort DH, Chia G, Treff NR, et al. Human embryos commonly form abnormal nuclei during development: a mechanism of DNA damage, embryonic aneuploidy, and developmental arrest. Hum Reprod 2016;31:312-23. [PubMed]
- van Echten-Arends J, Mastenbroek S, Sikkema-Raddatz B, et al. Chromosomal mosaicism in human preimplantation embryos: a systematic review. Hum Reprod Update 2011;17:620-7. [Crossref] [PubMed]
- Rubio C, Rodrigo L, Mercader A, et al. Impact of chromosomal abnormalities on preimplantation embryo development. Prenat Diagn 2007;27:748-56. [Crossref] [PubMed]
- Liu J, Wang W, Sun X, et al. DNA microarray reveals that high proportions of human blastocysts from women of advanced maternal age are aneuploid and mosaic. Biol Reprod 2012;87:148. [Crossref] [PubMed]
- Coonen E, Derhaag JG, Dumoulin JC, et al. Anaphase lagging mainly explains chromosomal mosaicism in human preimplantation embryos. Hum Reprod 2004;19:316-24. [Crossref] [PubMed]
- Daphnis DD, Delhanty JD, Jerkovic S, et al. Detailed FISH analysis of day 5 human embryos reveals the mechanisms leading to mosaic aneuploidy. Hum Reprod 2005;20:129-37. [Crossref] [PubMed]
- Gonzalez-Merino E, Emiliani S, Vassart G, et al. Incidence of chromosomal mosaicism in human embryos at different developmental stages analyzed by fluorescence in situ hybridization. Genet Test 2003;7:85-95. [Crossref] [PubMed]
- Munné S, Velilla E, Colls P, et al. Self-correction of chromosomally abnormal embryos in culture and implications for stem cell production. Fertil Steril 2005;84:1328-34. [Crossref] [PubMed]
- Sundvall L, Lund H, Niemann I, et al. Tetraploidy in hydatidiform moles. Hum Reprod 2013;28:2010-20. [Crossref] [PubMed]
- Decordier I, Cundari E, Kirsch-Volders M. Mitotic checkpoints and the maintenance of the chromosome karyotype. Mutat Res 2008;651:3-13. [Crossref] [PubMed]
- Kiessling AA, Bletsa R, Desmarais B, et al. Genome-wide microarray evidence that 8-cell human blastomeres over-express cell cycle drivers and under-express checkpoints. J Assist Reprod Genet 2010;27:265-76. [Crossref] [PubMed]
- Musacchio A, Salmon ED. The spindle-assembly checkpoint in space and time. Nat Rev Mol Cell Biol 2007;8:379-93. [Crossref] [PubMed]
- Nilsson J, Yekezare M, Minshull J, et al. The APC/C maintains the spindle assembly checkpoint by targeting Cdc20 for destruction. Nat Cell Biol 2008;10:1411-20. [Crossref] [PubMed]
- Los FJ, Van Opstal D, van den Berg C. The development of cytogenetically normal, abnormal and mosaic embryos: a theoretical model. Hum Reprod Update 2004;10:79-94. [Crossref] [PubMed]
- Harrison RH, Kuo HC, Scriven PN, et al. Lack of cell cycle checkpoints in human cleavage stage embryos revealed by a clonal pattern of chromosomal mosaicism analysed by sequential multicolour FISH. Zygote 2000;8:217-24. [Crossref] [PubMed]
- Michaelis C, Ciosk R, Nasmyth K. Cohesins: chromosomal proteins that prevent premature separation of sister chromatids. Cell 1997;91:35-45. [Crossref] [PubMed]
- Brooker AS, Berkowitz KM. The roles of cohesins in mitosis, meiosis, and human health and disease. Methods Mol Biol 2014;1170:229-66. [Crossref] [PubMed]
- Mantikou E, Wong KM, Repping S, et al. Molecular origin of mitotic aneuploidies in preimplantation embryos. Biochim Biophys Acta 2012;1822:1921-30. [Crossref] [PubMed]
- Solomon DA, Kim T, Diaz-Martinez LA, et al. Mutational inactivation of STAG2 causes aneuploidy in human cancer. Science 2011;333:1039-43. [Crossref] [PubMed]
- Zhang N, Ge G, Meyer R, et al. Overexpression of Separase induces aneuploidy and mammary tumorigenesis. Proc Natl Acad Sci U S A 2008;105:13033-8. [Crossref] [PubMed]
- Singh VP, Gerton JL. Cohesin and human disease: lessons from mouse models. Curr Opin Cell Biol 2015;37:9-17. [Crossref] [PubMed]
- Yanez LZ, Han J, Behr BB, et al. Human oocyte developmental potential is predicted by mechanical properties within hours after fertilization. Nat Commun 2016;7:10809. [Crossref] [PubMed]
- Silber S, Escudero T, Lenahan K, et al. Chromosomal abnormalities in embryos derived from testicular sperm extraction. Fertil Steril 2003;79:30-8. [Crossref] [PubMed]
- García-Ferreyra J, Luna D, Villegas L, et al. High Aneuploidy Rates Observed in Embryos Derived from Donated Oocytes are Related to Male Aging and High Percentages of Sperm DNA Fragmentation. Clin Med Insights Reprod Health 2015;9:21-7. [Crossref] [PubMed]
Cite this article as: Stolakis V, Bertero MC. Molecular aspects of aneuploidy in preimplantation human embryos: a mini-review. Ann Res Hosp 2019;3:8.


